Data Content in CPRCSdb
CPRCSdb is a curated database that links tumor single-cell heterogeneity to key clinical phenotypes. It includes data from approximately 4.05 million cells across 1,053 single-cell samples and 101 datasets, alongside 11,032 bulk RNA-seq samples, covering 29 cancer types.
The database provides comprehensive information on five clinical phenotypes: patient survival, tumor stage, size, lymph node involvement, and distant metastasis, along with 30 anticancer drug response phenotypes. Users can perform fuzzy matching searches for scRNA and bulk RNA data, explore drug-cancer associations through dual search paths, and utilize six enrichment analysis tools related to cancer prognosis.
Detailed sample pages offer insights into differential expression, enrichment analysis, and cell-cell communication, making CPRCSdb a valuable resource for investigating tumor heterogeneity and its impact on cancer progression and treatment outcomes.
How to Use CPRCSdb
CPRCSdb (http://www.yzbio.top/CPRCSdb/) is a comprehensive database that links tumor single-cell heterogeneity with key clinical phenotypes and drug responses. Here's how to get started:
- Search for Data: Use the search bar on the homepage to perform fuzzy matching searches by sample ID, cancer type, or tissue name.
- Browse Single-cell Data: Explore over 4.05 million cells across 1,053 single-cell samples and 101 datasets. View quality control metrics, differential expression, and cell-cell communication analysis.
- Explore Bulk RNA-seq Samples: Access 11,032 bulk RNA-seq samples covering 29 cancer types, annotated with five clinical phenotypes including survival, tumor stage, and metastasis.
- Investigate Drug Responses: Discover associations between 30 anti-cancer drugs and cancer projects using dual search paths (Drug → Project or Project → Drug).
- Analyze Results: Utilize six integrated enrichment tools and survival analysis modules to explore gene functions and prognosis-related features.
CPRCSdb provides an intuitive platform for researchers to investigate how tumor heterogeneity influences cancer progression and treatment outcomes.
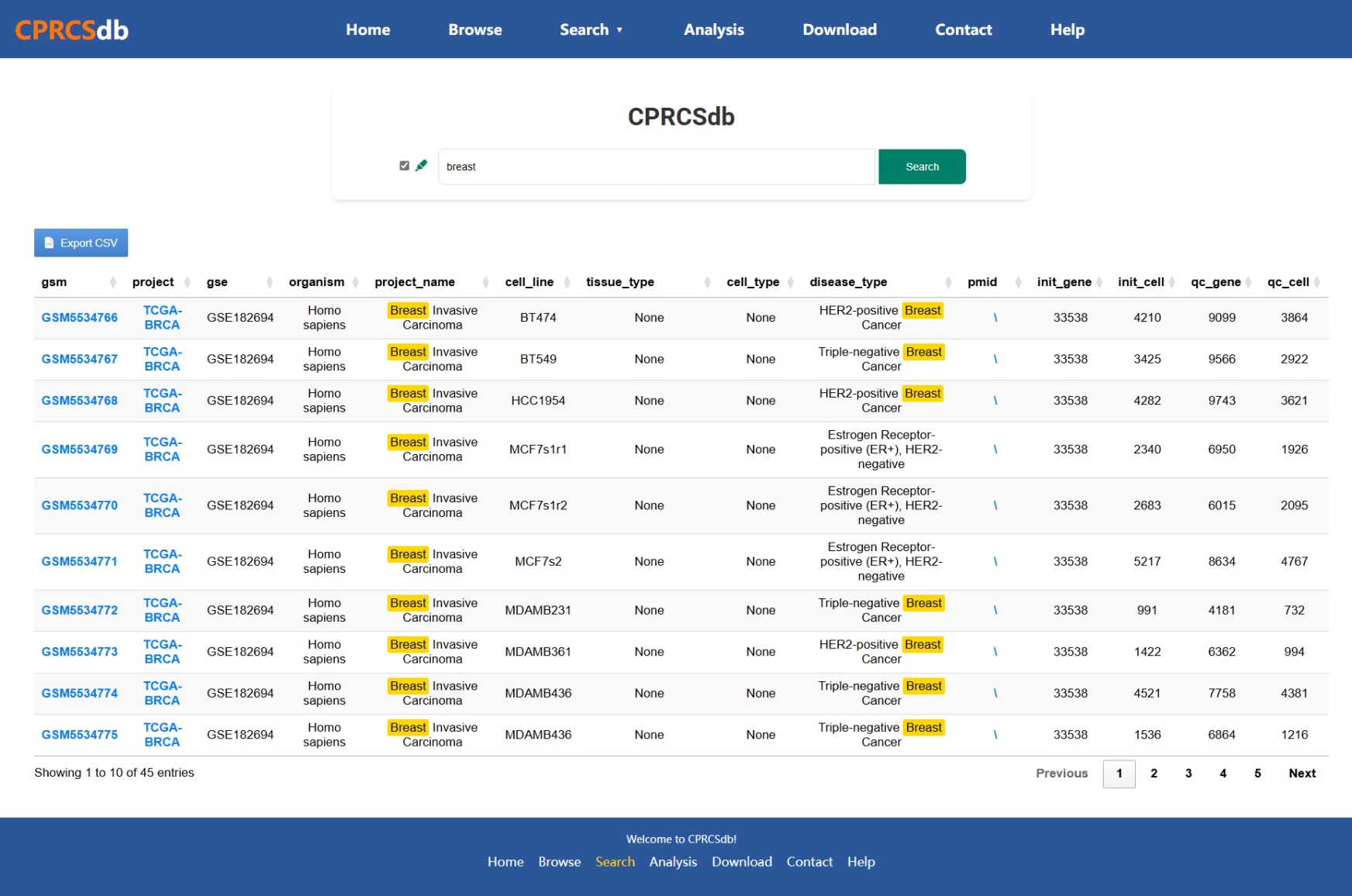
How to Search in CPRCSdb
Method 1: Fuzzy Matching Search for Single-cell Datasets and Bulk Projects
- Locate the Search Bar: On the homepage, find the search bar usually located at the top or center of the page.
- Enter Keywords: Input relevant keywords such as sample ID, cancer name, tissue name, etc.
- View Results: The search results will be displayed in a table format
including GSM ID, cancer name, QC status, etc.
- Click on the GSM ID to access detailed single-cell sample pages
with four sections:
- Sample Overview
- Bulk to Single-Cell Label Mapping
- Differential and Enrichment Analysis
- Cell-Cell Communication Analysis
- Click on the project abbreviation to view bulk sample detail pages
containing three sections:
- Bulk Sample Details
- Single-Cell Sample Information
- Differential Survival Analysis
- Click on the GSM ID to access detailed single-cell sample pages
with four sections:
Method 2: Exploring Drug-Cancer Associations through Dual Search Paths
Given the complex many-to-many relationship between drugs and cancer projects, we define two paths for locating drug phenotype annotations:
- Drug → Project: Select a drug to see which specific cancer projects it applies to.
- Project → Drug: Select a cancer project to view the anticancer drugs used.
On the drug-focused detail page, you will find basic drug information, single-cell sample details, and drug response analysis results.

Data Browsing Guide
Below are example screenshots of the CPRCSdb interface to help you better understand how to browse and search for different types of data.
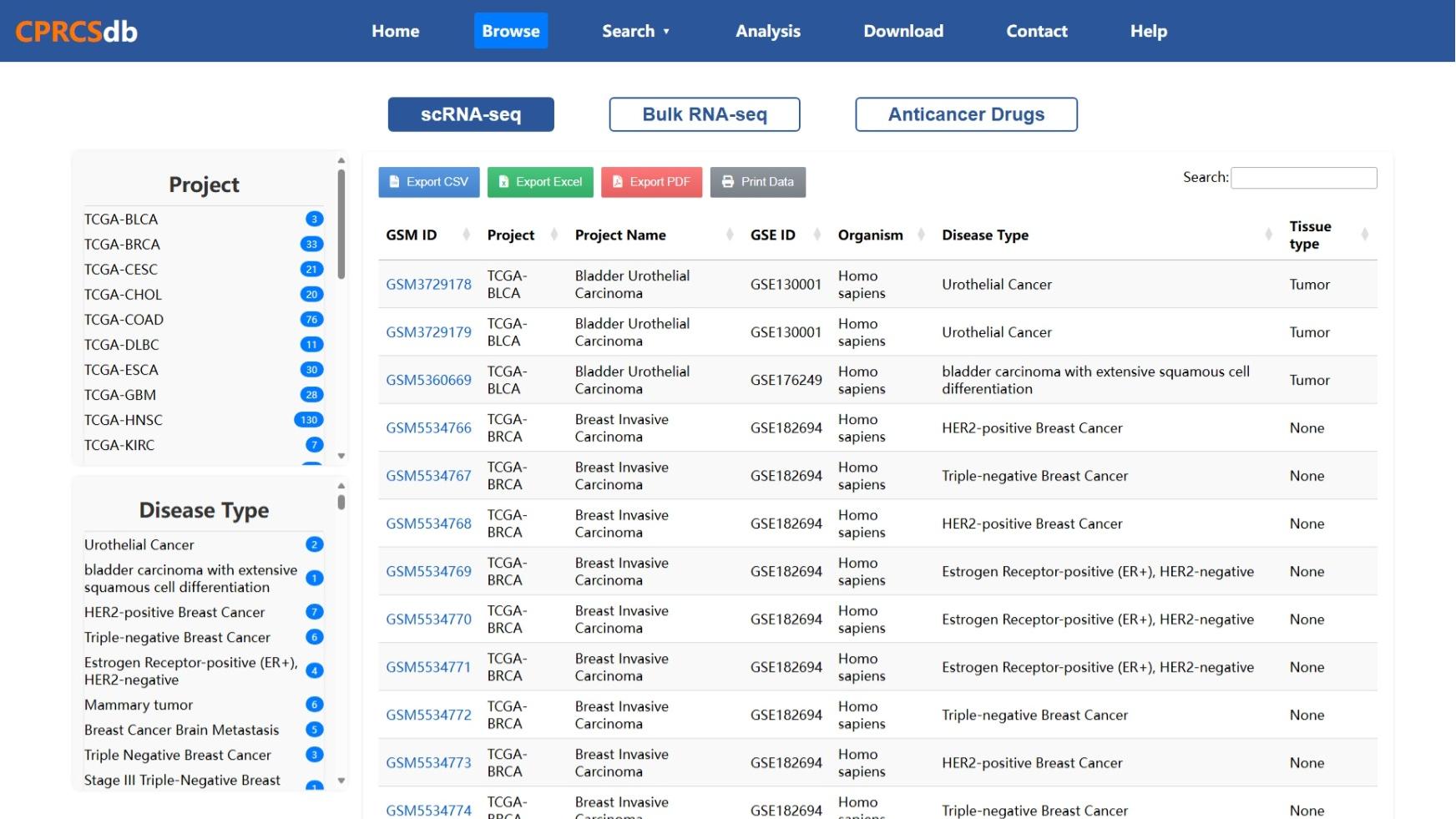
Figure 1: Single-cell Data Browsing Interface
This image shows the interface for browsing single-cell RNA-seq datasets in CPRCSdb. You can search or filter by cancer type, tissue source, or experimental platform. Each entry includes metadata such as GSM ID, sample description, and quality control status.
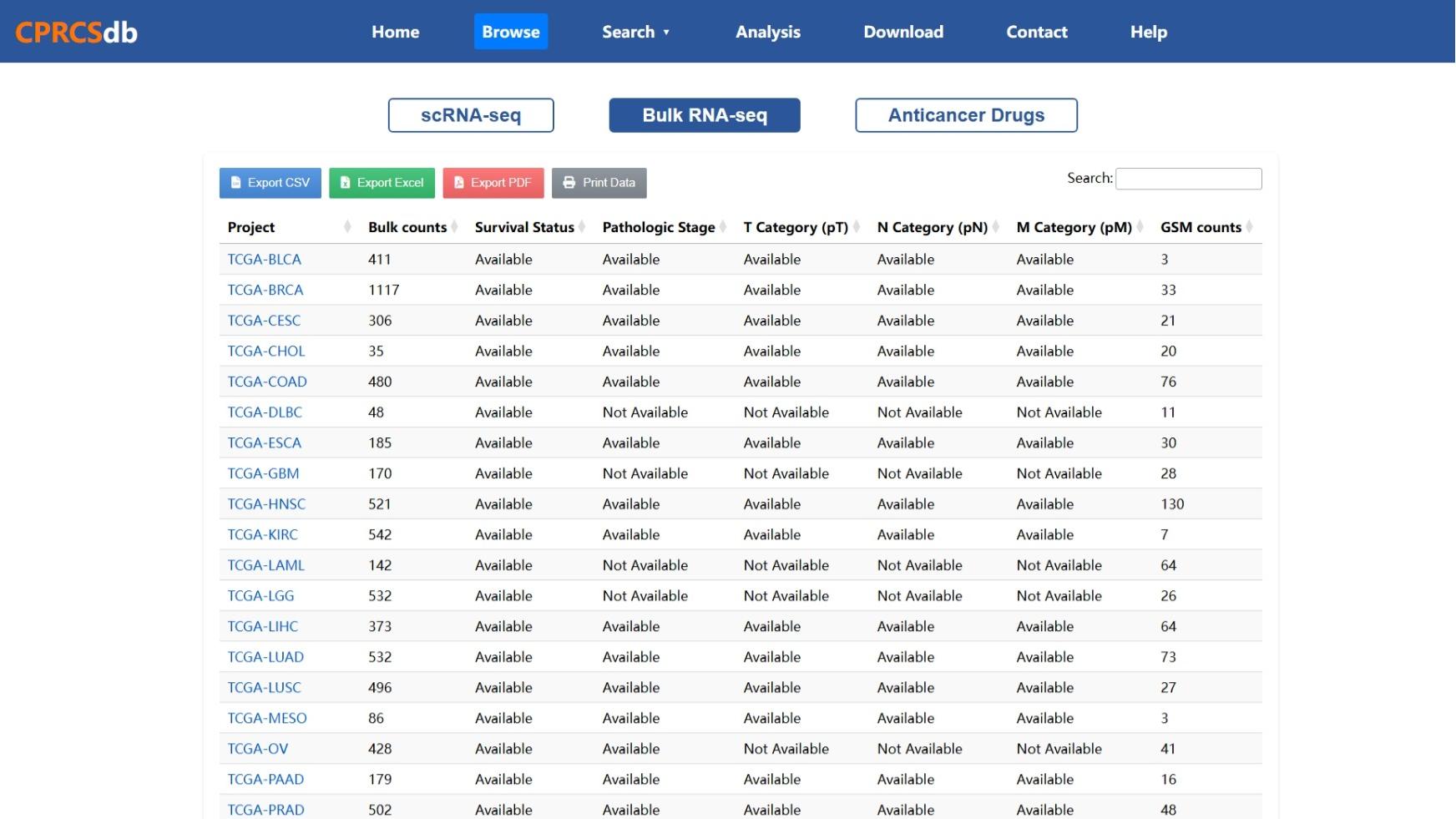
Figure 2: Bulk RNA-seq Data Browsing Interface
This screenshot demonstrates how bulk RNA-seq samples are displayed in CPRCSdb. It includes clinical phenotypes such as tumor stage, survival status, lymph node involvement, and drug response.
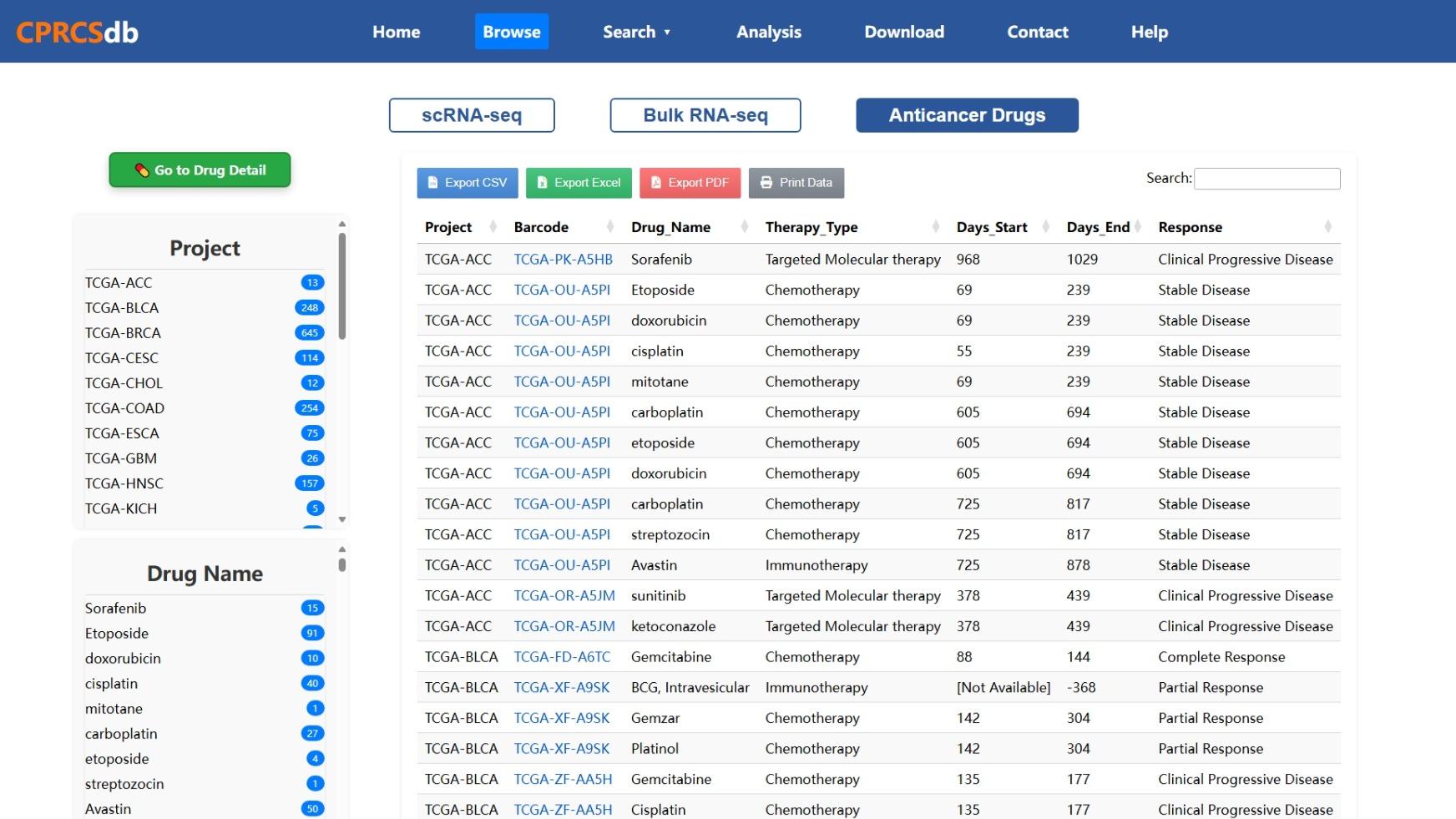
Figure 3: Anti-cancer Drug Response Interface
This image illustrates the drug response section in CPRCSdb. It displays associations between drugs and cancer projects, allowing users to explore drug sensitivity/resistance profiles and perform downstream analysis like enrichment and survival analysis.
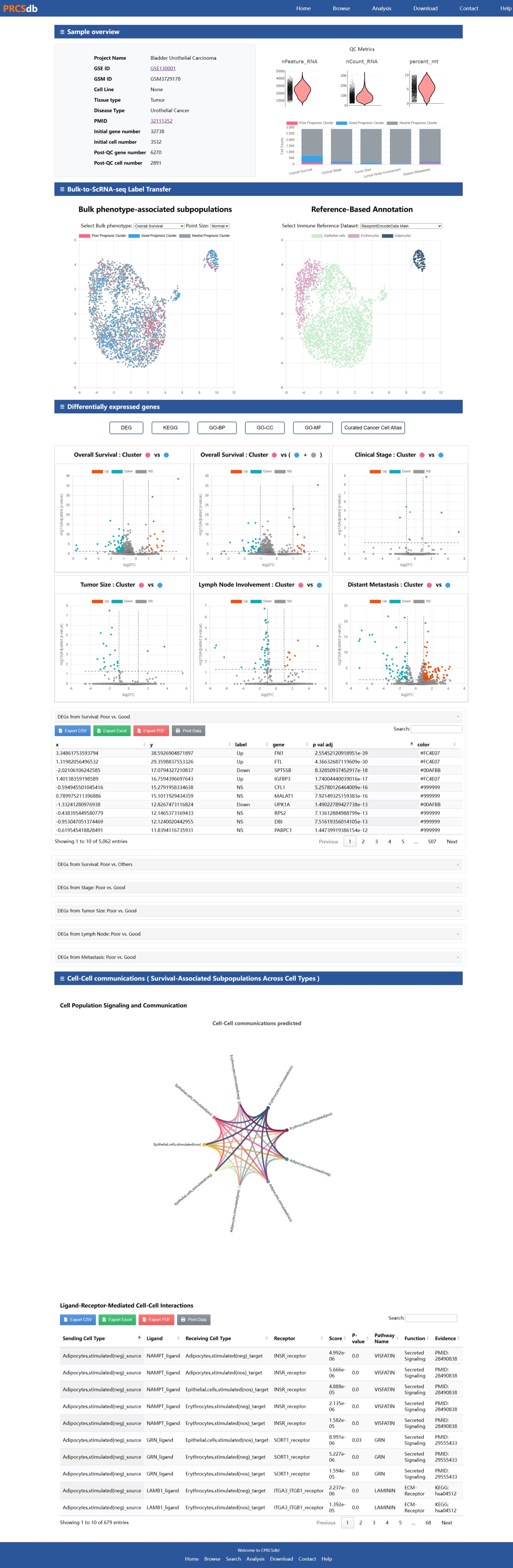
Single-cell Sample
Below is an example screenshot of the Single-cell Sample interface in CPRCSdb to assist you in navigating and utilizing this section effectively.
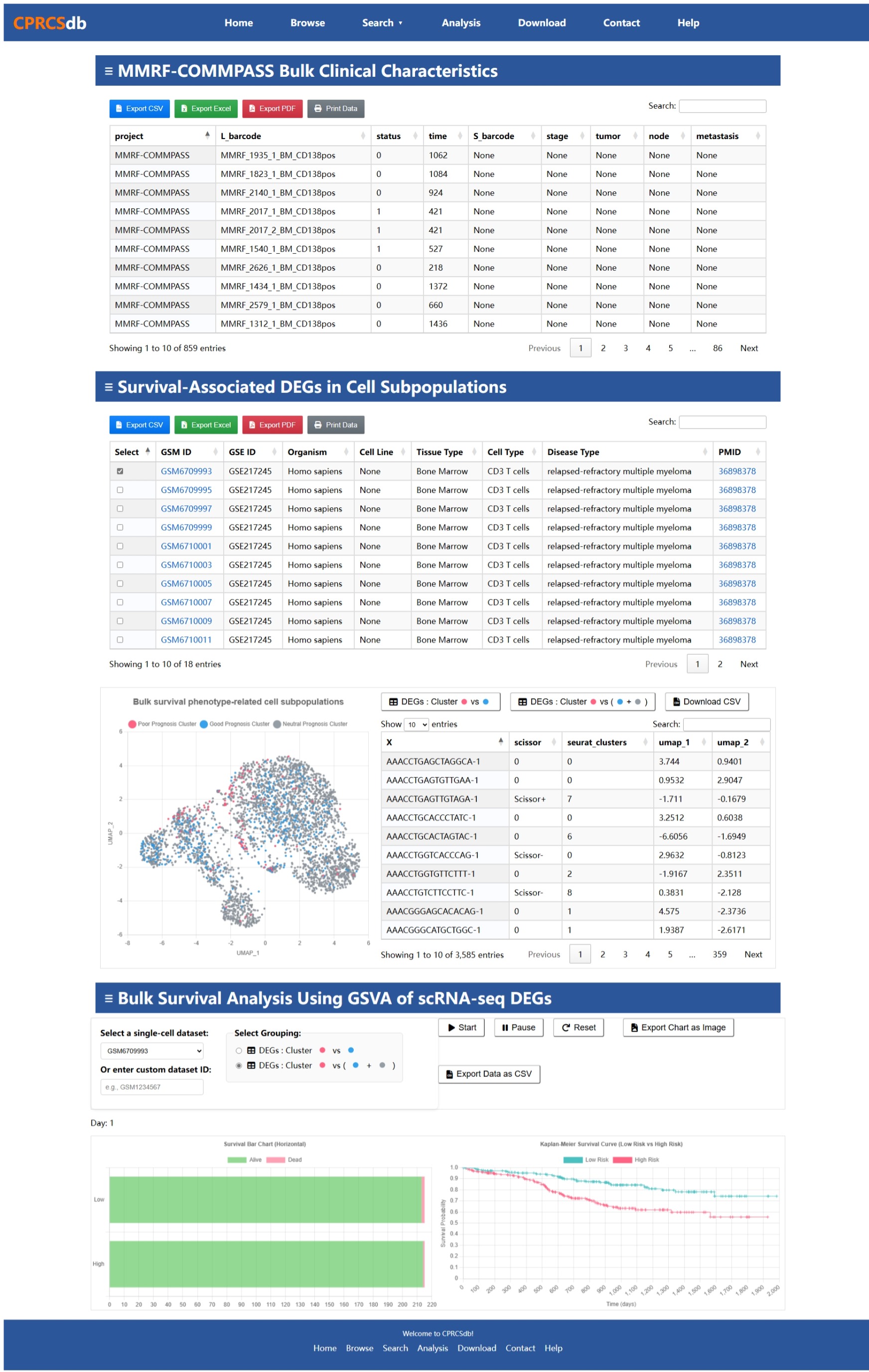
Bulk Project
Below is an example screenshot of the Bulk project interface in CPRCSdb to assist you in navigating and utilizing this section effectively.
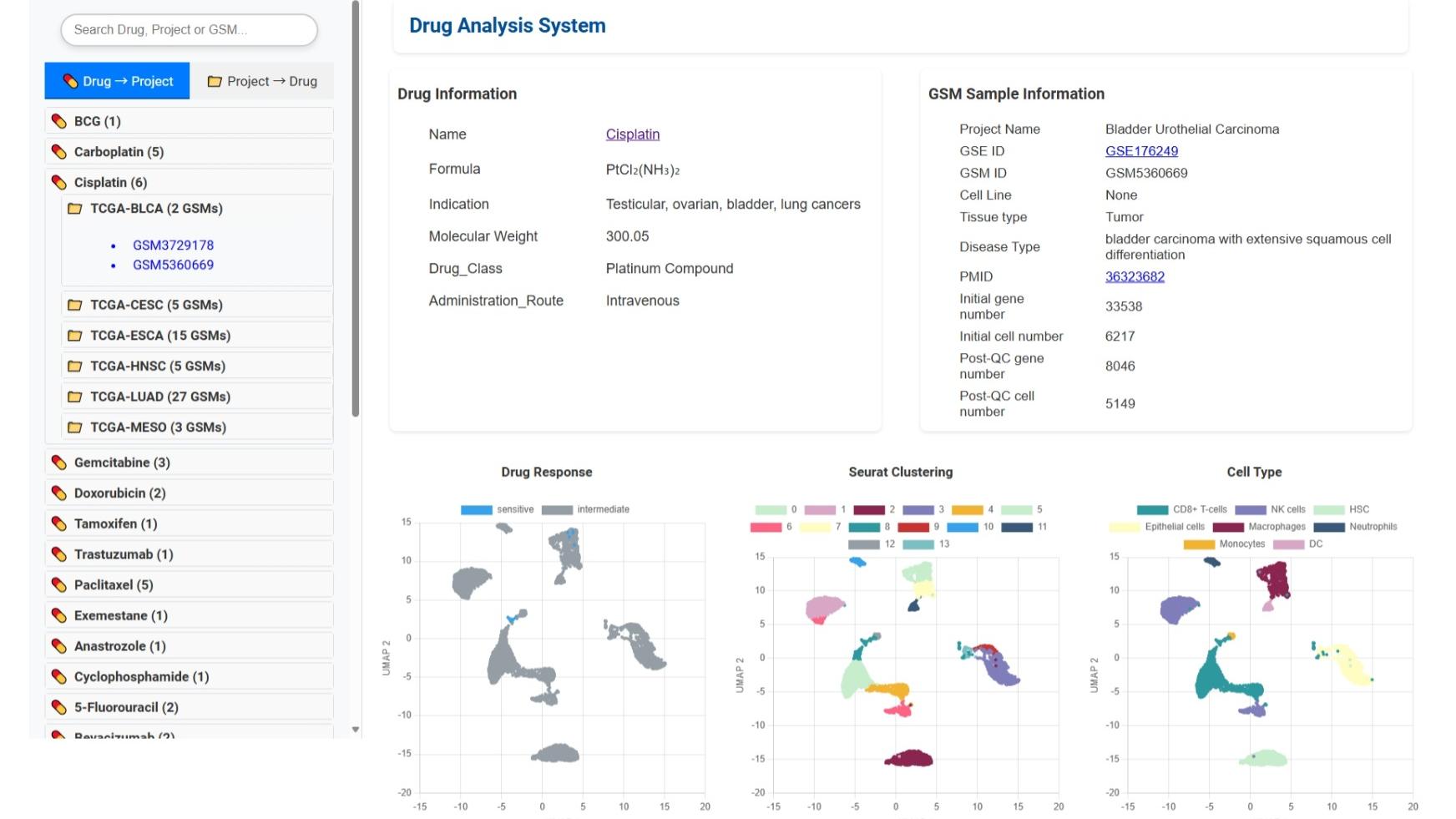
Drug Analysis System
Below is an example screenshot of the drug detail interface in CPRCSdb to help you understand how to interpret and use this section.
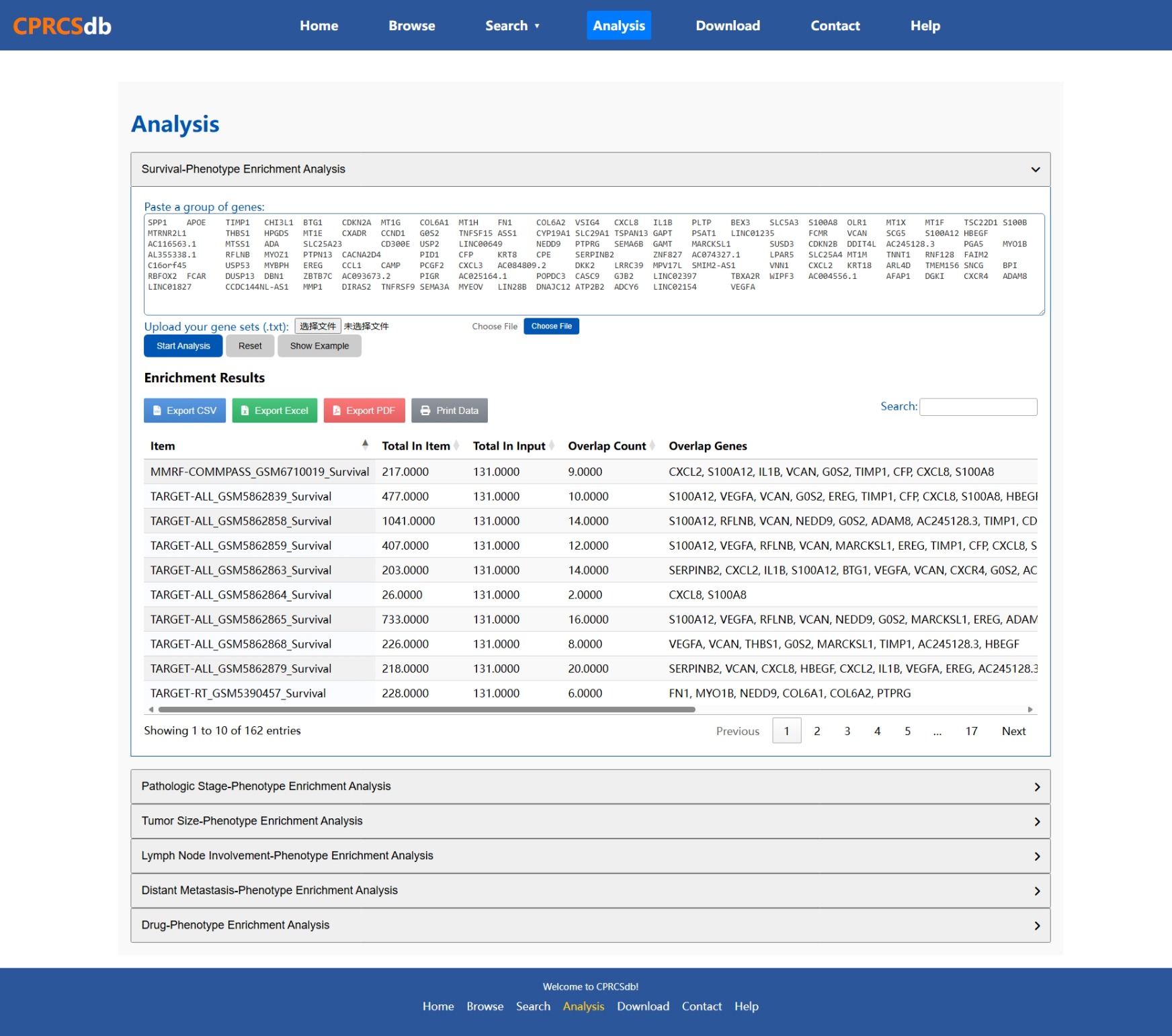
Analysis
Below is an example screenshot of the analysis interface in CPRCSdb.
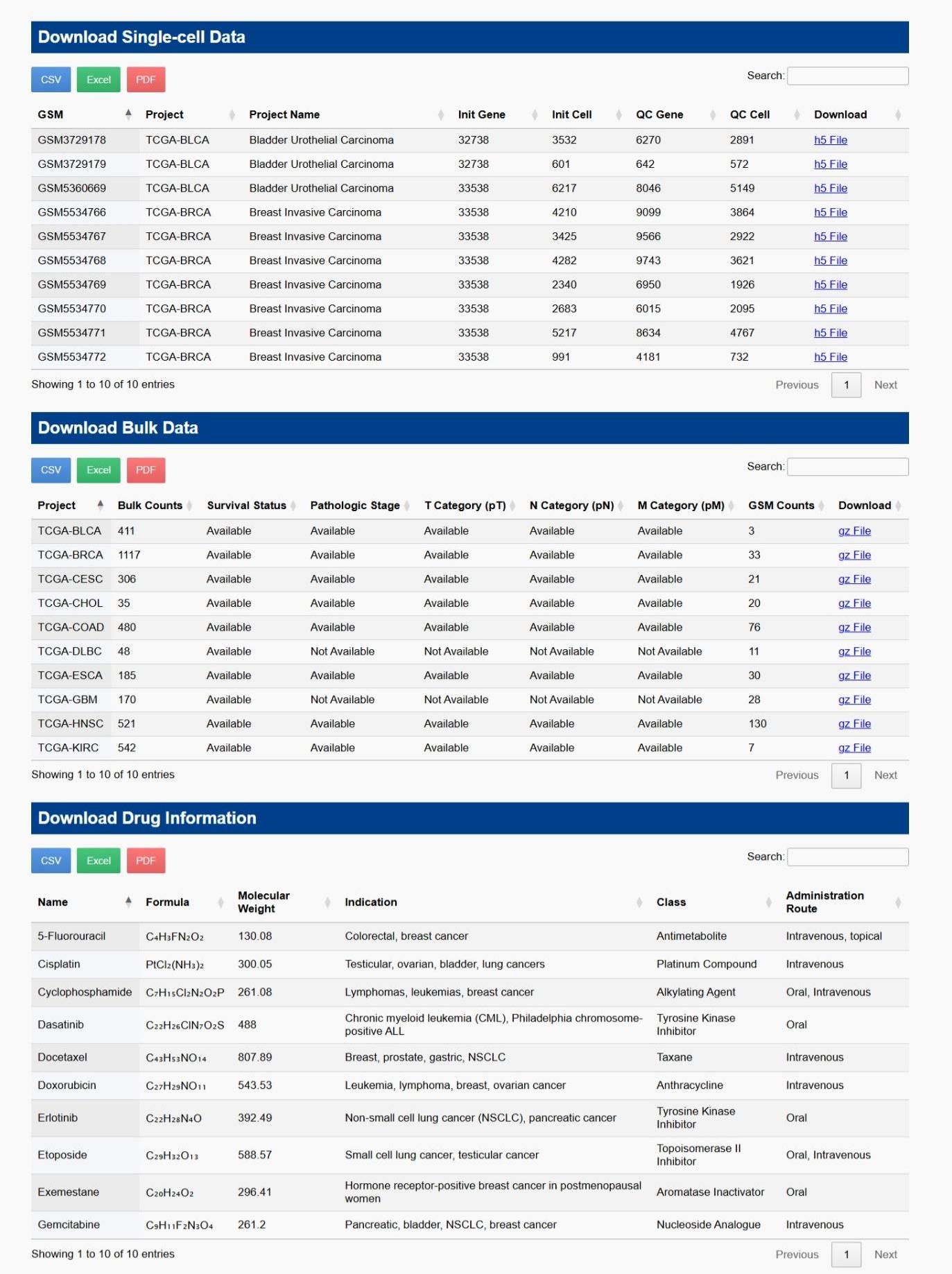
Download
Below is an example screenshot of the download interface in CPRCSdb.