Welcome to CPRCSdb!
What is CPRCSdb?
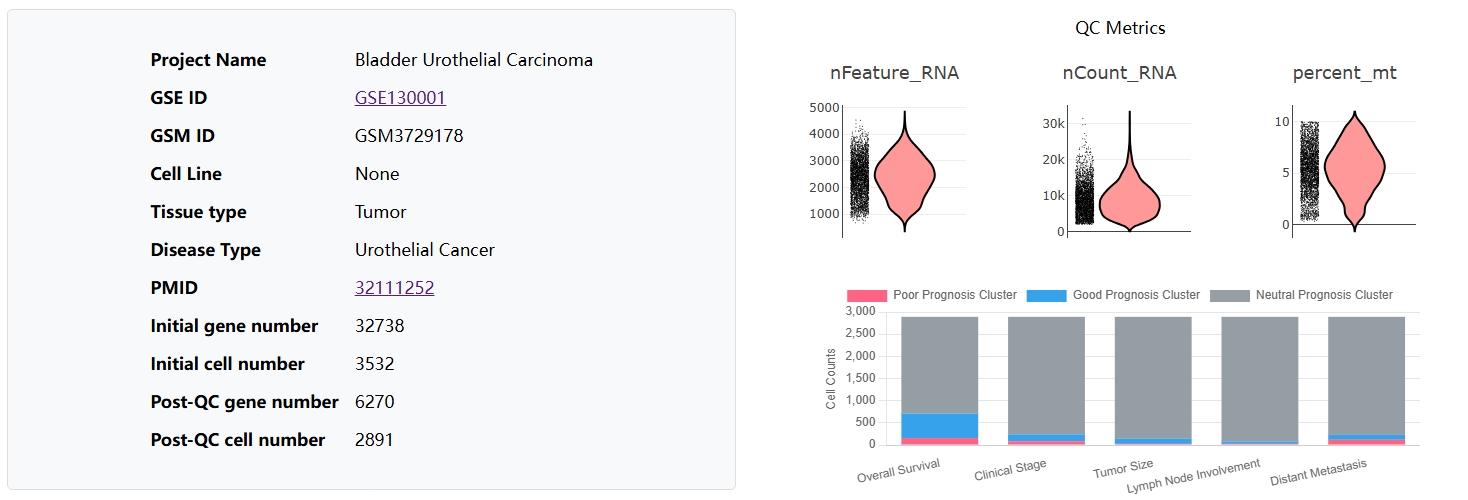
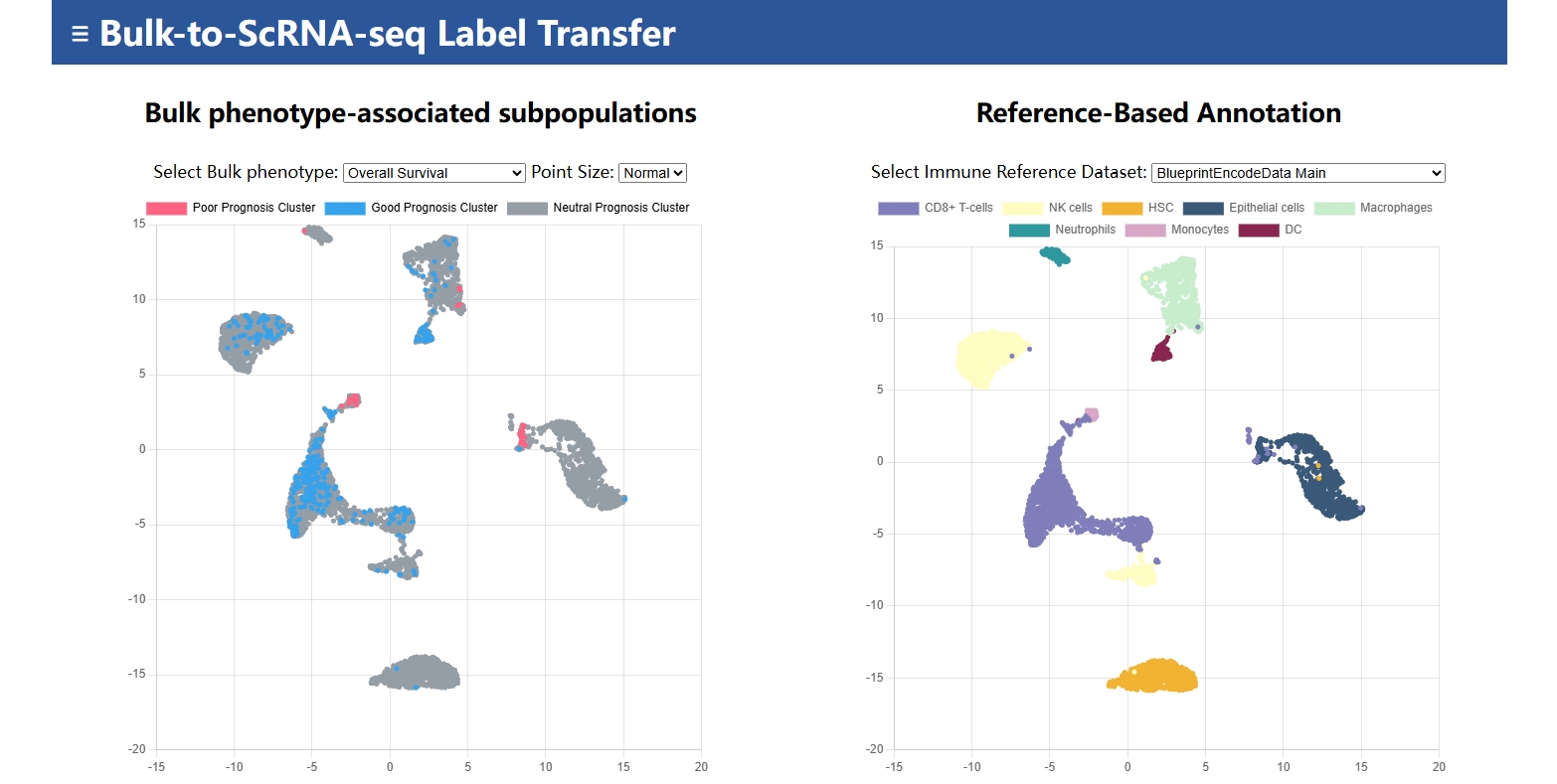
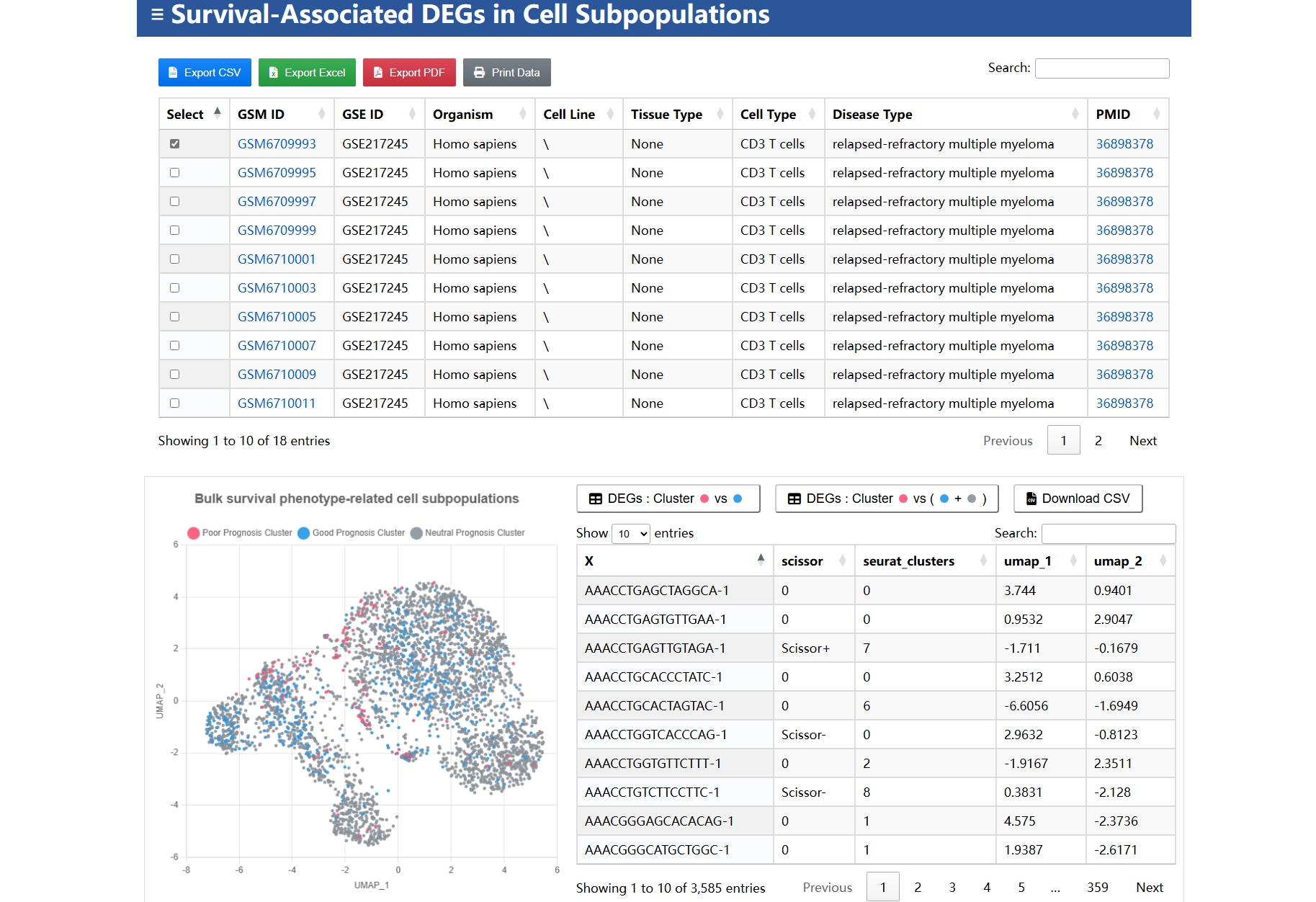
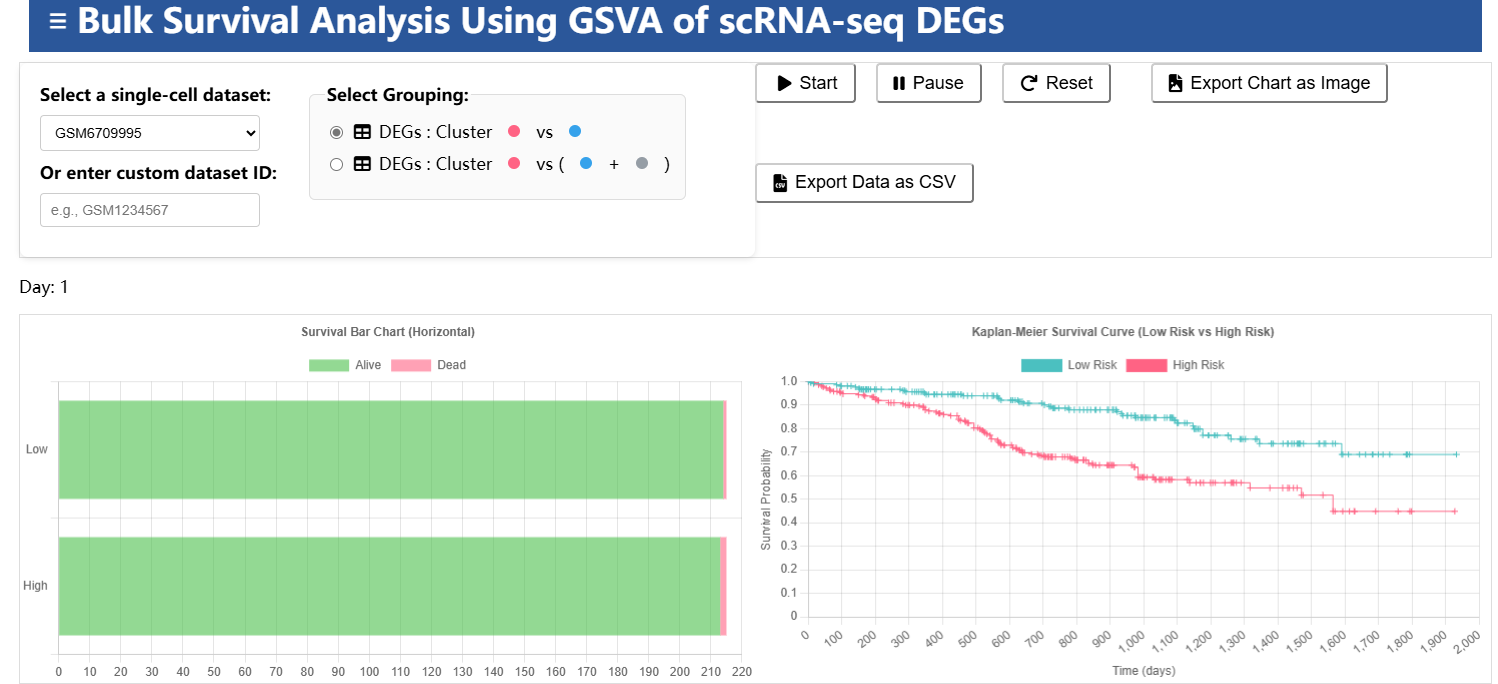
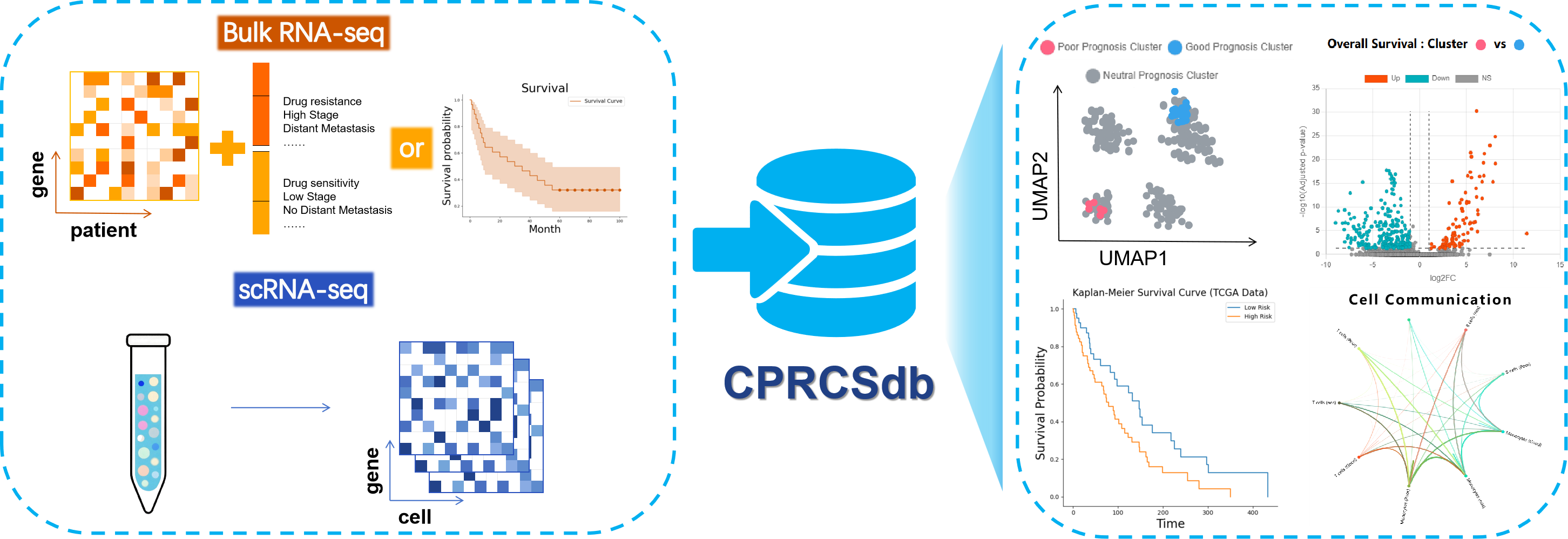
Cancer-Phenotype-Related Cancer Cell Subpopulations Database (CPRCSdb) is a database designed to systematically study how tumor single-cell heterogeneity influences tumor staging, patient survival, and responses to anti-cancer therapies. It employs Scissor, an algorithm published in Nature Biotechnology, to perform joint analysis. This approach preserves the rich clinical phenotype information from bulk data while capturing the cellular heterogeneity provided by single-cell data. The platform provides comprehensive bioinformatics analyses to explore the link between single-cell features and clinical phenotypes.
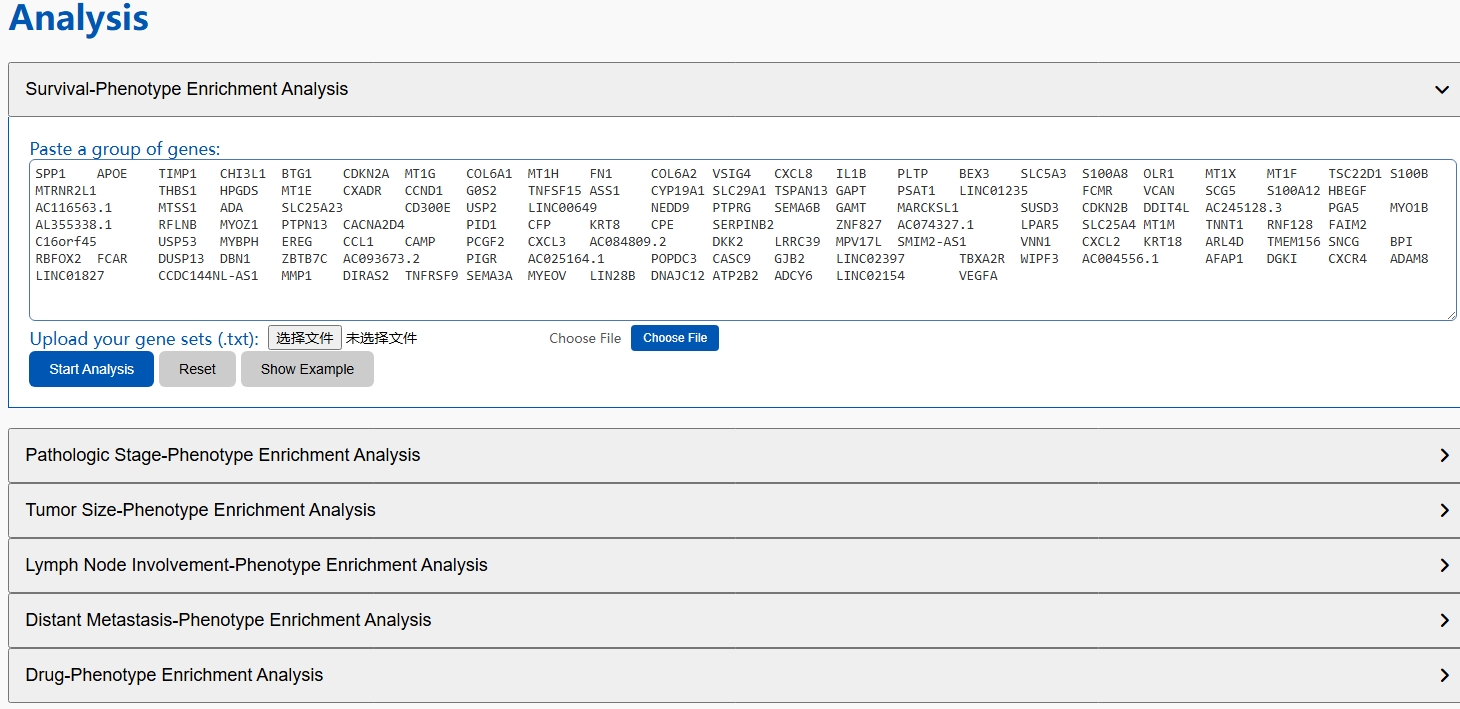
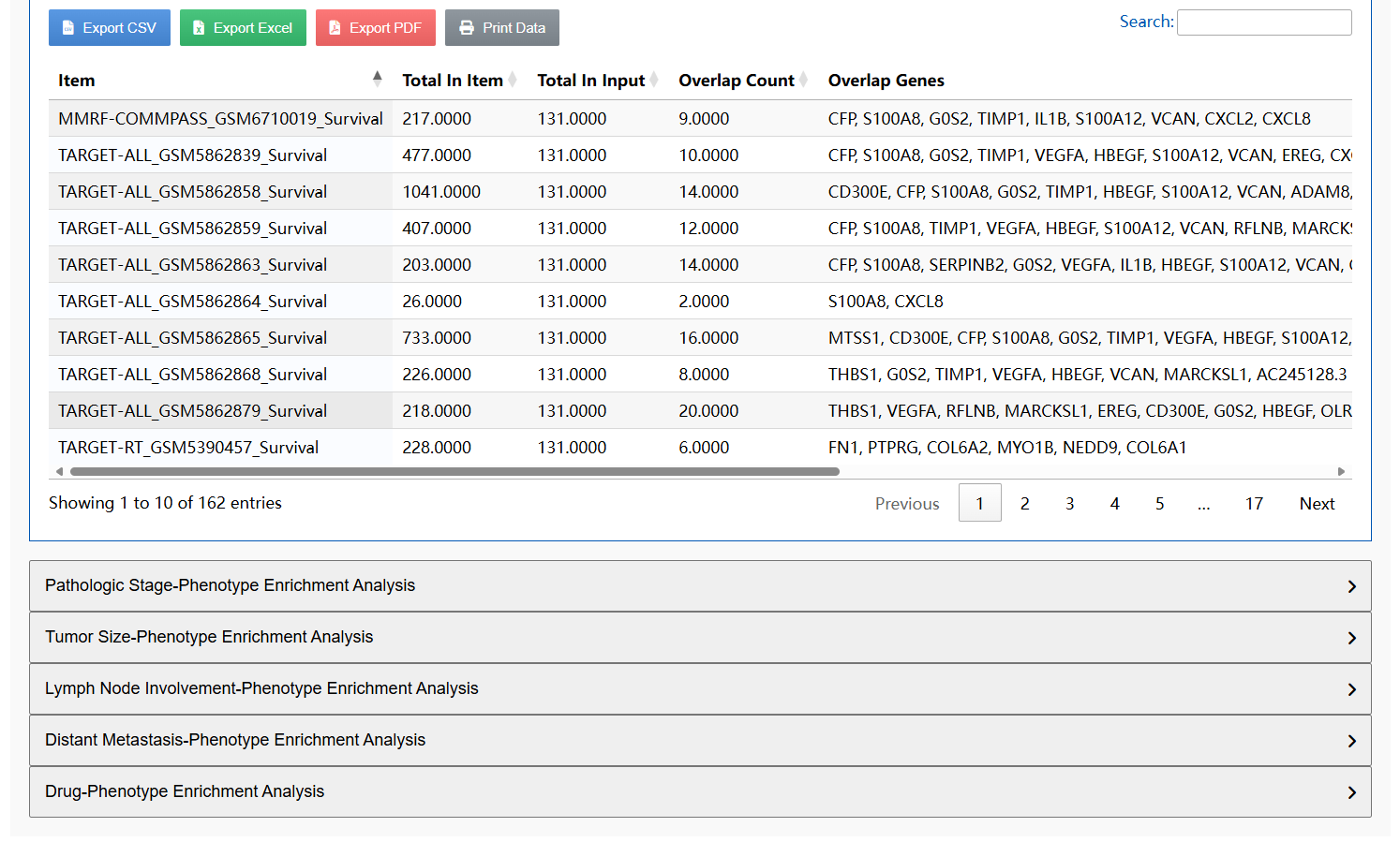
In brief, CPRCSdb curates data from over 4.05 million cells derived from approximately 1,053 manually collected single-cell samples and 101 integrated datasets, along with 11,032 bulk RNA-seq samples, collectively covering 29 cancer types. These bulk samples cover five clinical phenotypes, namely patient survival status, tumor stage, tumor size, lymph node involvement, and distant metastasis, as well as 30 anti-cancer drug response phenotypes, each supported by at least three sensitive and three resistant samples.
🔍 Diverse Search Options for Precise Exploration
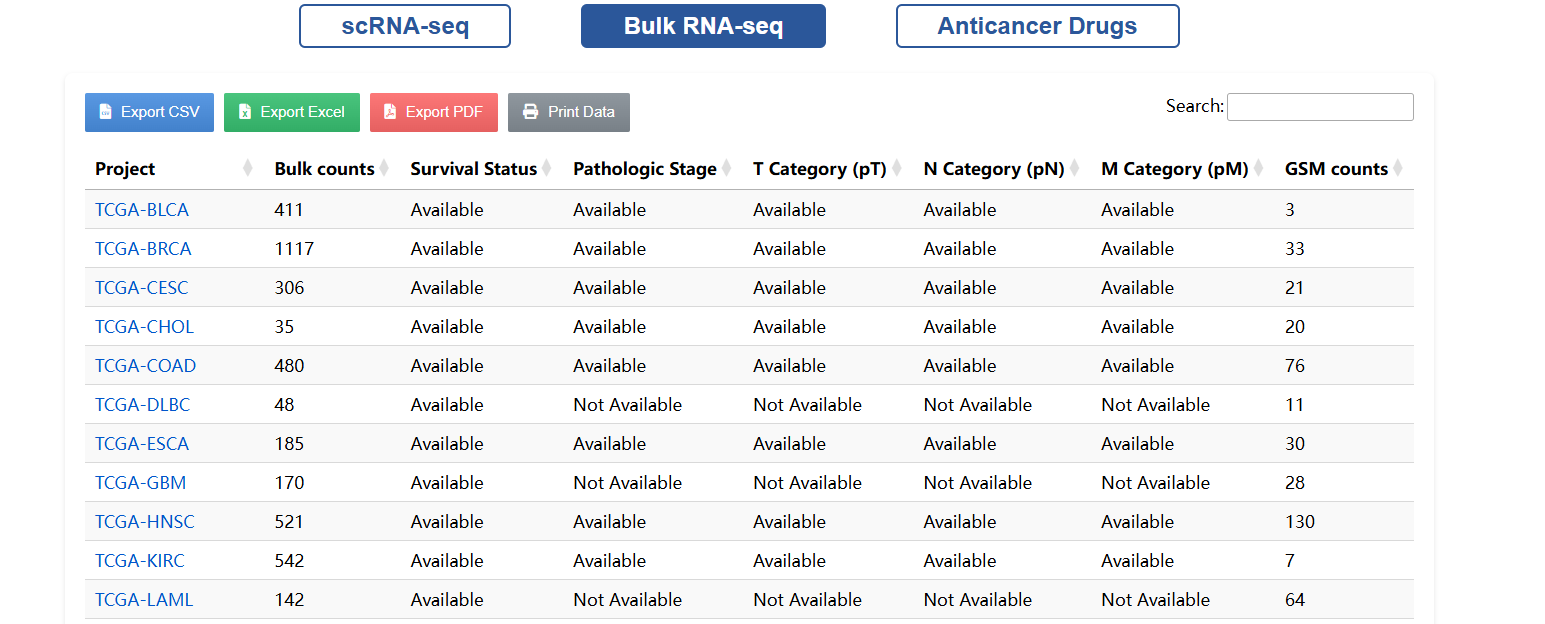
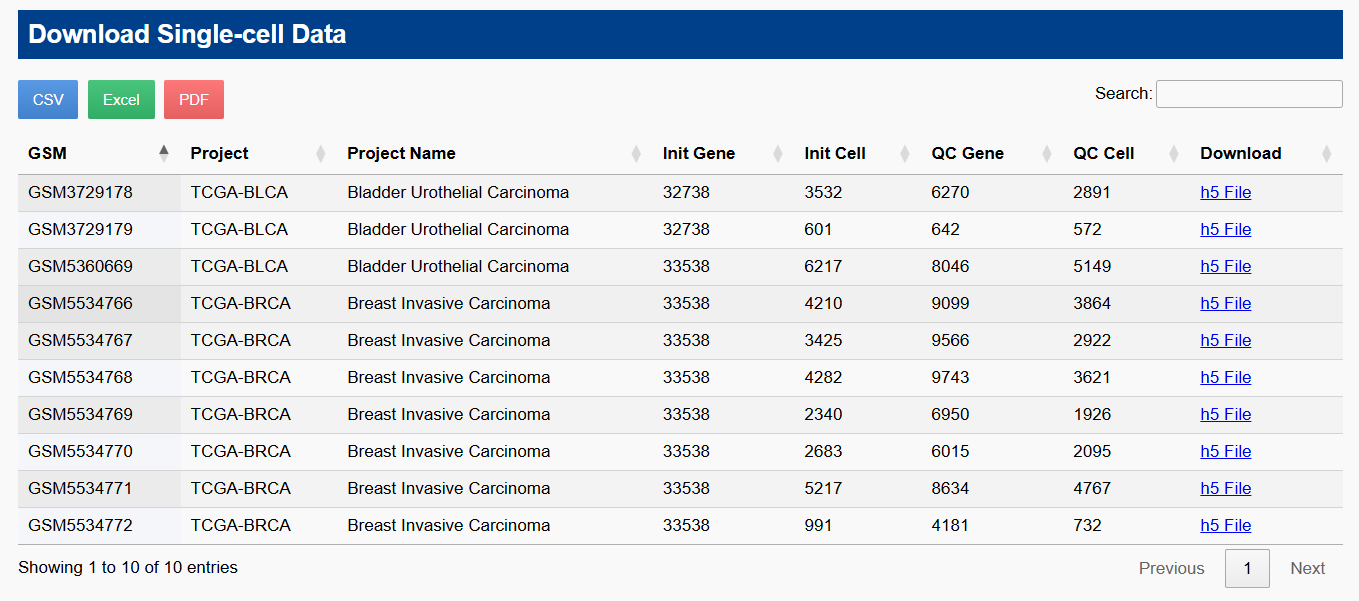
🔬 Search Single-cell & Bulk RNA Data
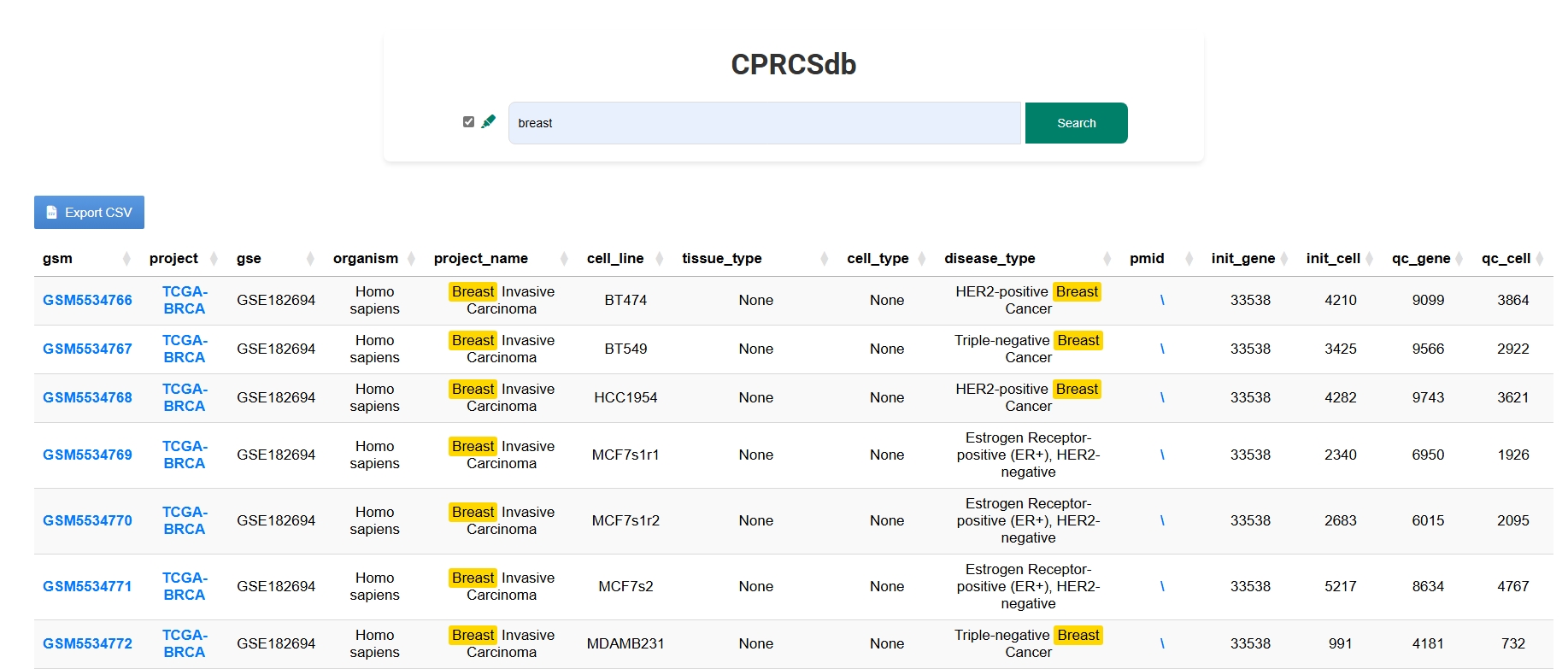
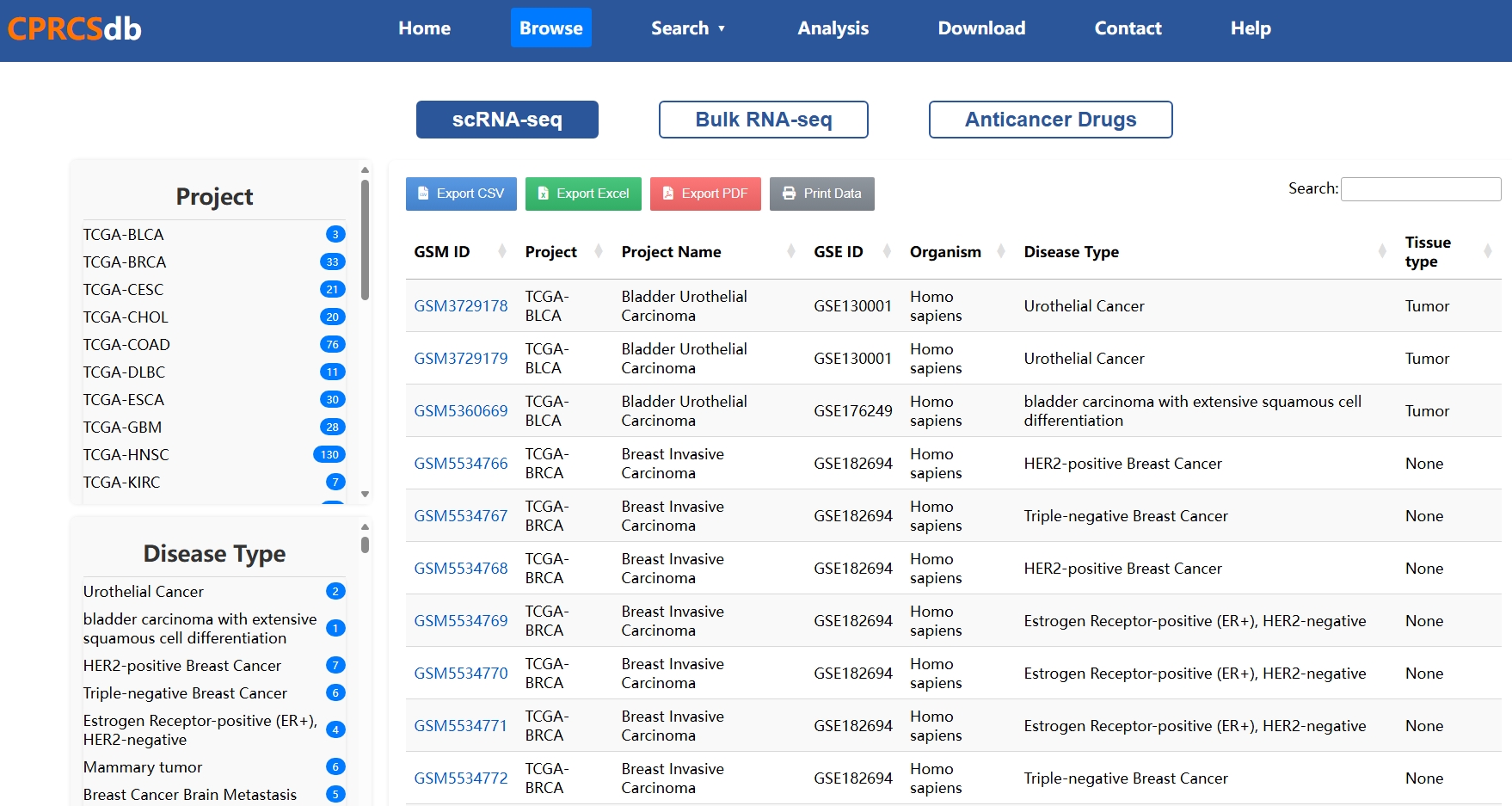
Use keywords like sample ID, cancer type, or tissue name to perform fuzzy matching searches. Results are clearly summarized, with direct links to detailed single-cell profiles or bulk sample stratification.
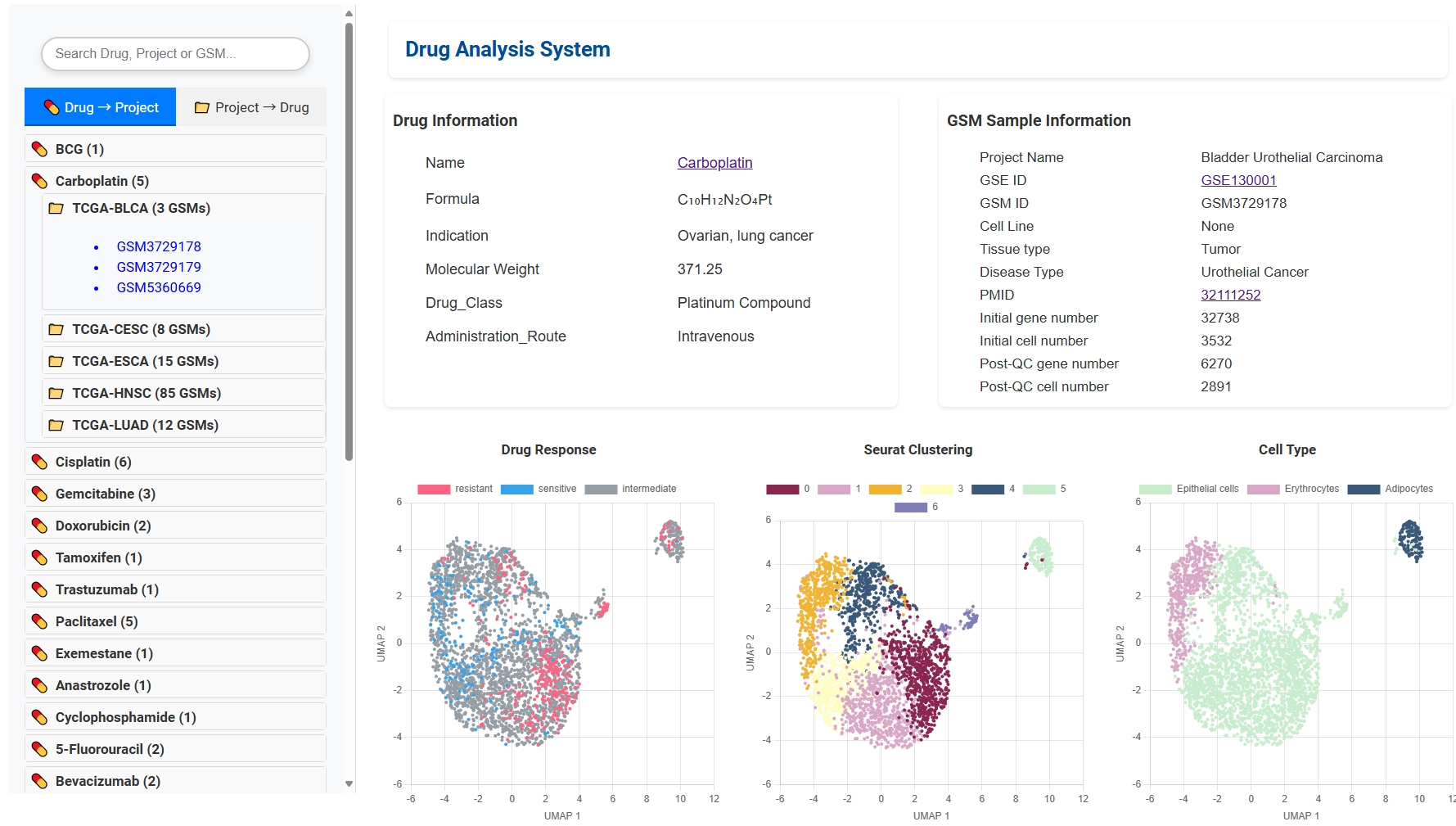
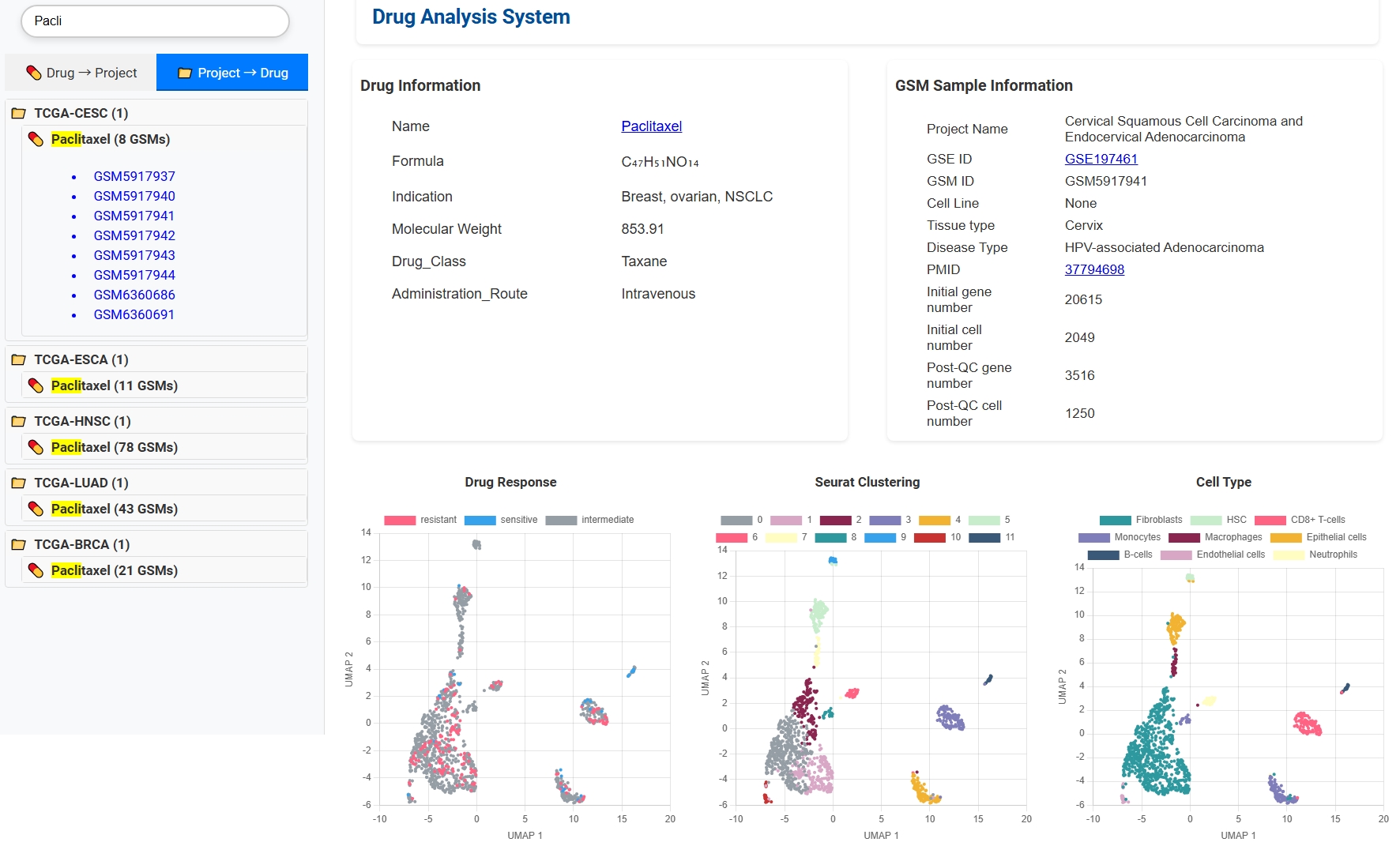
🔄 Bidirectional Drug-Cancer Search
Explore drugs by cancer project or find which cancers a specific drug is used for. Drug detail pages provide essential drug info, associated single-cell samples, and response analysis results.
Data Analysis Tools and Reference Datasets
| Tool / Dataset | Full Name / Source | Function / Description |
|---|---|---|
Scissor(2.1.0) |
R package | Mapping of the "bulk" label to scRNA. |
SingleR(2.4.1) |
R package | Automated Cell-Type Annotation |
TCGAbiolinks(2.30.4) |
R package | Download TCGA RNA-seq and clinical data. |
BlueprintEncodeData.rds |
BLUEPRINT & ENCODE Project | Gene expression profiles of various immune cell types, especially for hematopoietic studies. |
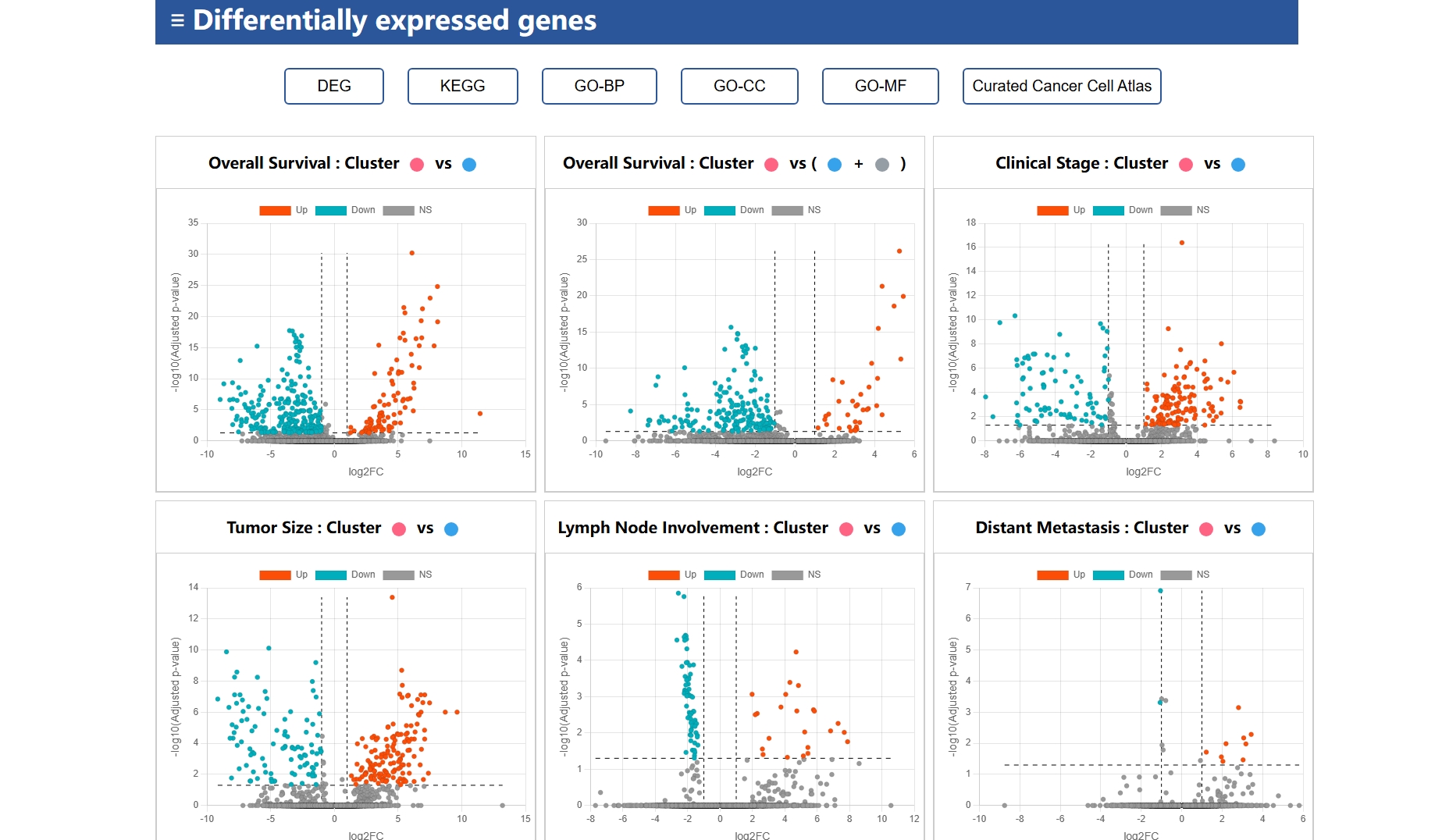
Seurat(5.0.3)::FindMarkers |
Seurat Function (R) | Identifies differentially expressed genes between clusters or conditions in scRNA-seq data. |
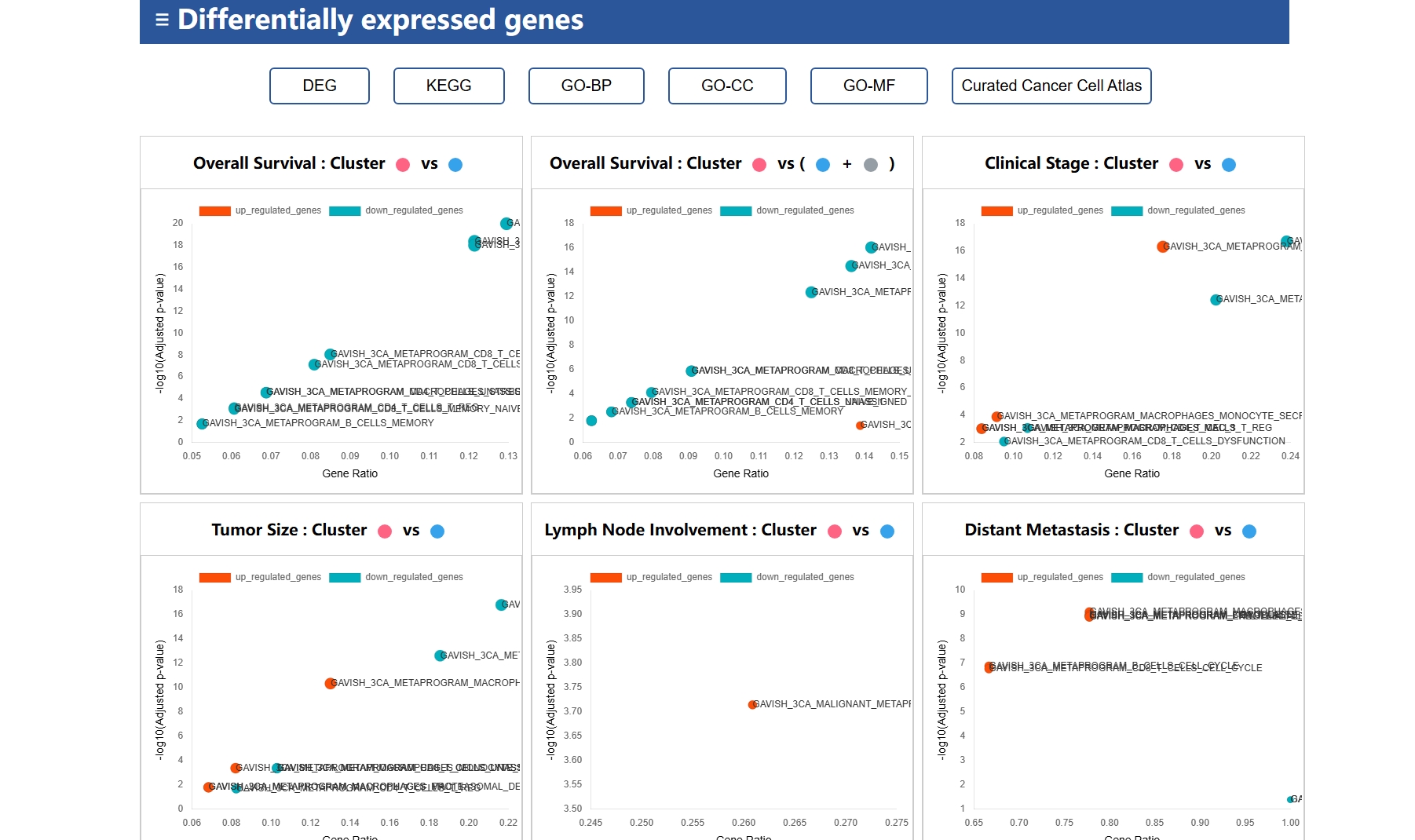
clusterProfiler(4.10.1) |
R package | Functional enrichment analysis for GO, KEGG, etc. |
GSVA(1.50.5) |
Gene Set Variation Analysis (R) | Score the samples using gene sets. |
CellChat(1.6.1) |
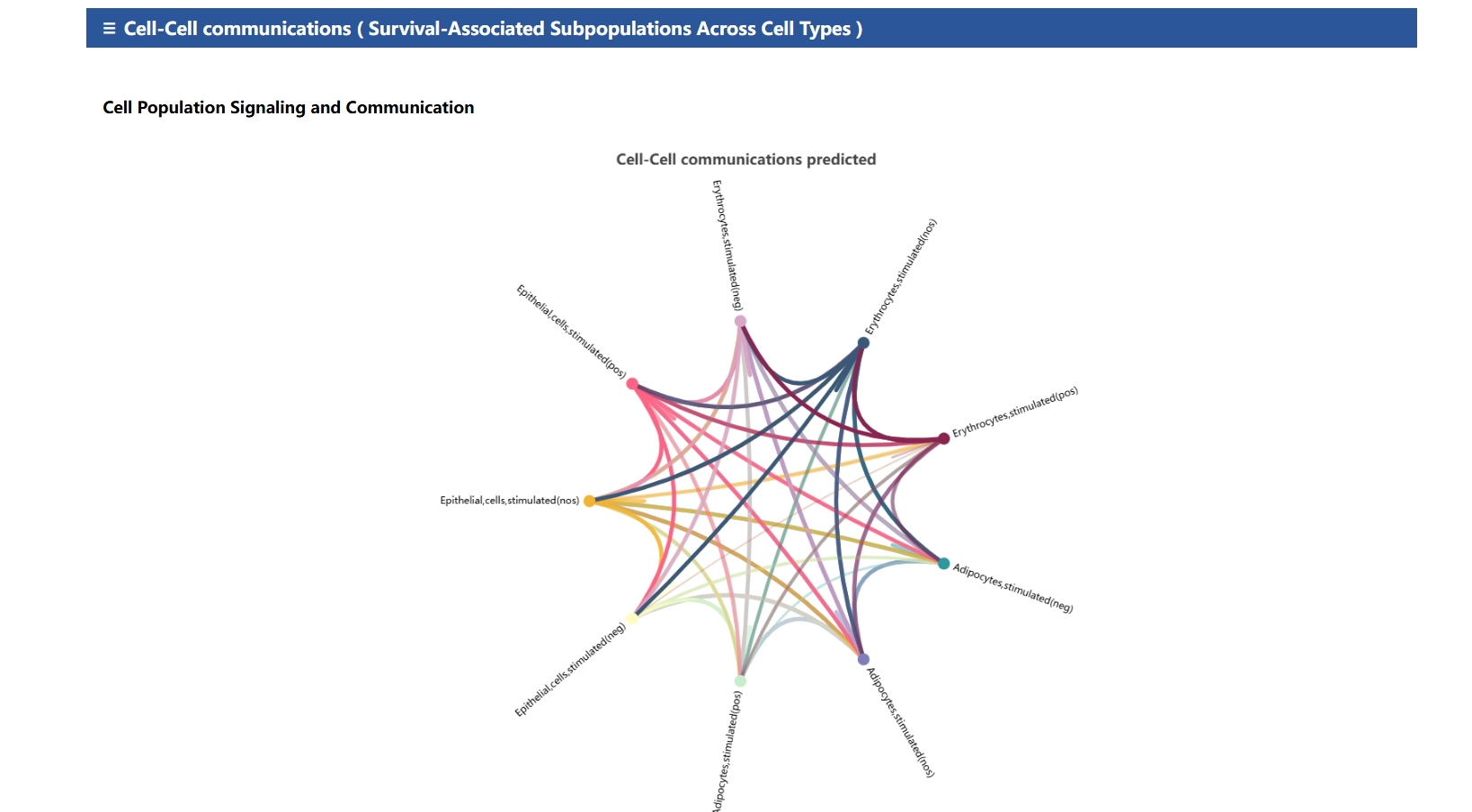
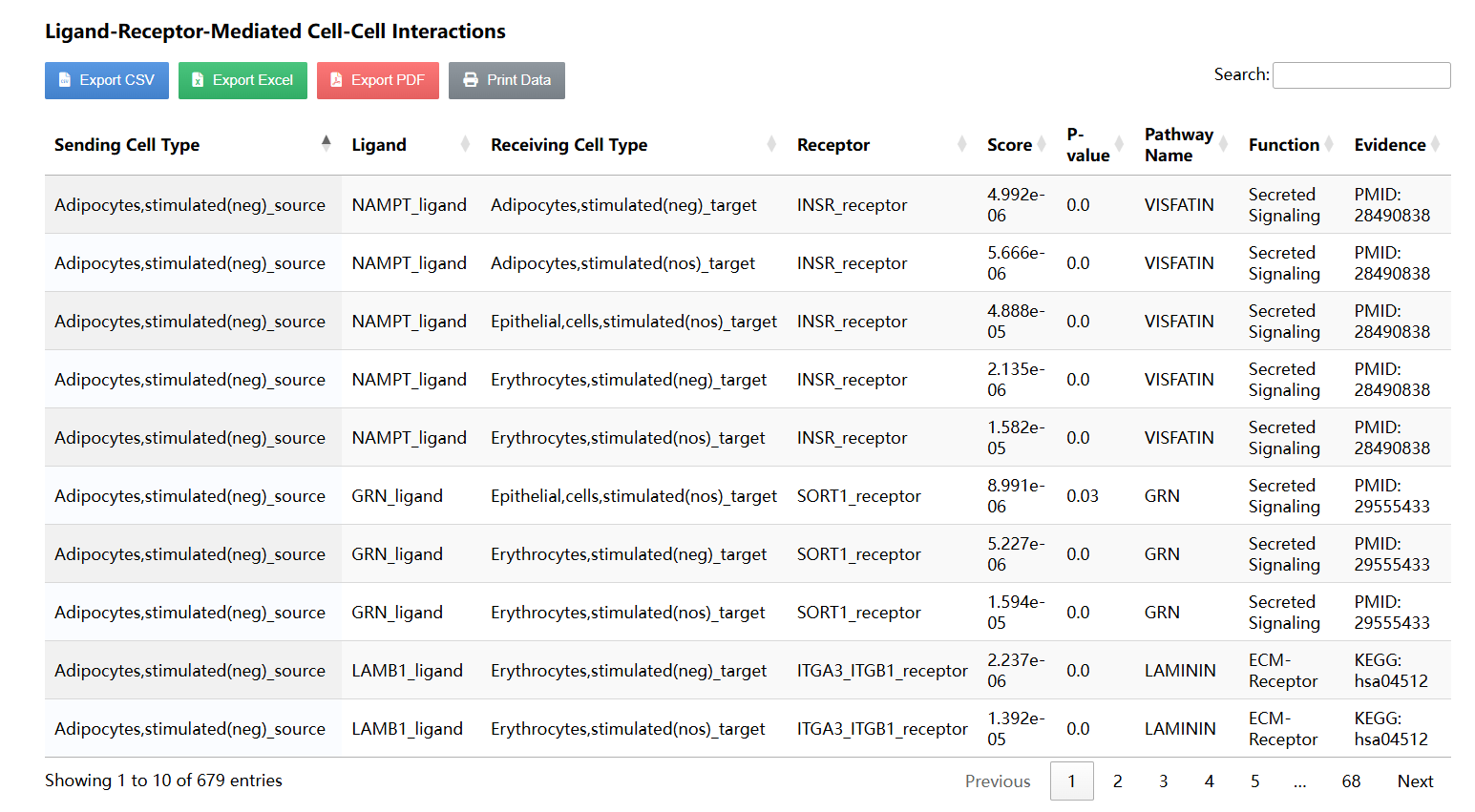
R package | Predicts intercellular communication networks based on ligand-receptor interactions. |
survival(3.8.3) & survminer(0.5.1) |
R packages | Performs Kaplan-Meier survival analysis and Cox regression modeling. |
Contact us
Principal Investigator: Desi Shang, Ph.D.
Institutional Address: Insititute of Biochemistry and Molecular Biology, Hengyang Medical College, University of South China, Hengyang, Hunan 421001, China
Email: sds_46@163.com
Fax: +86 0734 8279018
ORCID: 0000-0002-9141-393X